Quickstart¶
Import the common constructors:
from mcerp import *
Create Distributions¶
Construct uncertain variables from probability distributions:
x1 = N(24, 1) # normal distribution: mean 24, standard deviation 1
x2 = N(37, 4) # normal distribution: mean 37, standard deviation 4
x3 = Exp(2) # exponential distribution: lambda 2
The first four moments are available as properties:
x1.mean
x1.var
x1.std
x1.skew
x1.kurt
x1.stats
Calculate With Uncertainty¶
Use normal Python arithmetic:
Z = (x1 * x2**2) / (15 * (1.5 + x3))
Print the compact representation:
Z
uv(1161.35518507, 116688.945979, 0.353867228823, 3.00238273799)
Or print a labeled description:
Z.describe()
MCERP Uncertain Value:
> Mean................... 1161.35518507
> Variance............... 116688.945979
> Skewness Coefficient... 0.353867228823
> Kurtosis Coefficient... 3.00238273799
The values above may vary slightly between sessions because new samples are generated when variables are created.
Mathematical Functions¶
Use mcerp.umath for uncertainty-aware mathematical functions:
import mcerp.umath as umath
from mcerp import N
H = N(64, 0.5)
M = N(16, 0.1)
P = N(361, 2)
t = N(165, 0.5)
C = 38.4
Q = C * umath.sqrt((520 * H * P) / (M * (t + 460)))
Q.describe()
Probabilities¶
Comparison operators estimate sampled probabilities when an uncertain value is compared with a scalar:
x1 < 21
Z >= 1000
0.0014
0.6622
Discrete distributions can be queried with equality:
h = H(50, 5, 10)
h == 4
h <= 3
0.004
0.9959
For continuous distributions, exact equality usually returns zero probability:
n = N(0, 1)
n == 0
n < 0
n < 1.5
0.0
0.5
0.9332
When two uncertain values are compared, mcerp performs a paired t-test on the
underlying samples to decide whether one distribution is statistically greater
or less than the other.
rvs1 = N(5, 10)
rvs2 = N(5, 10) + N(0, 0.2)
rvs3 = N(8, 10) + N(0, 0.2)
rvs1 < rvs2
rvs1 < rvs3
False
True
Equality between two uncertain values checks whether they have the same root samples and therefore the same propagated moments:
Z * Z == Z**2
True
Plot Distributions¶
Plotting requires Matplotlib:
from mcerp import N
x = N(24, 1)
x.plot(show=True)
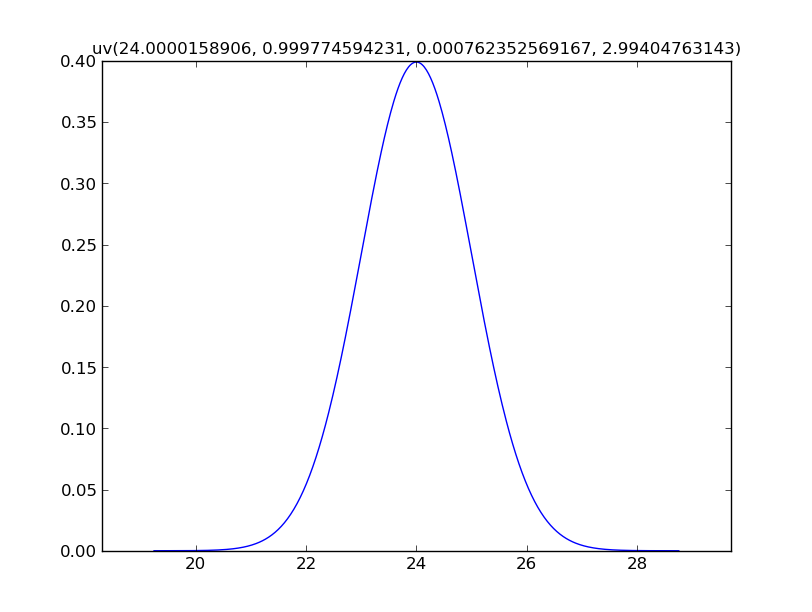
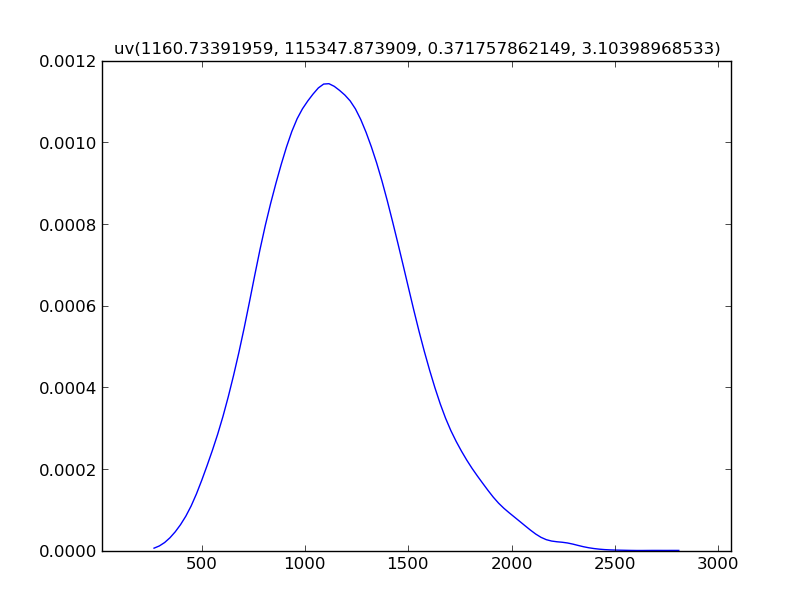
For calculated values, mcerp uses a kernel density estimate by default,
because a closed-form PDF or PMF may not be available:
Z.plot()
Z.show()
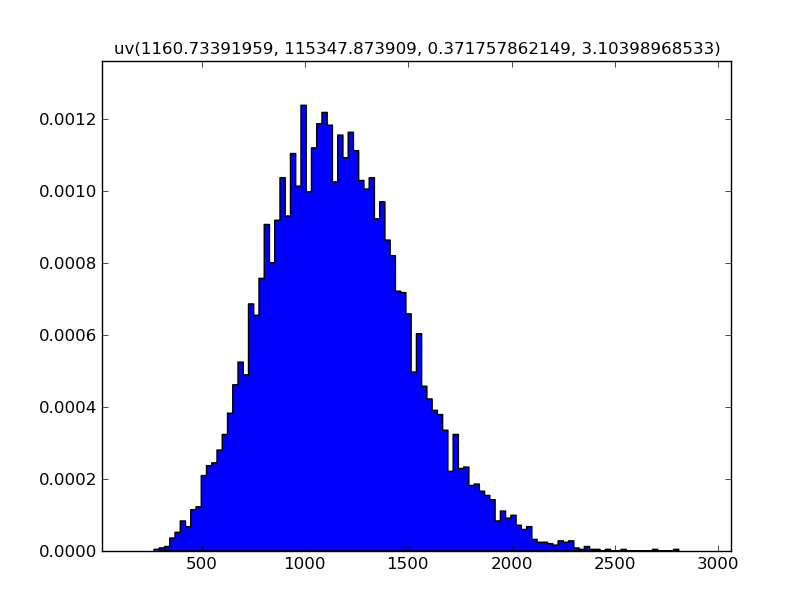
Use hist=True to show a histogram instead:
Z.plot(hist=True)
Z.show()
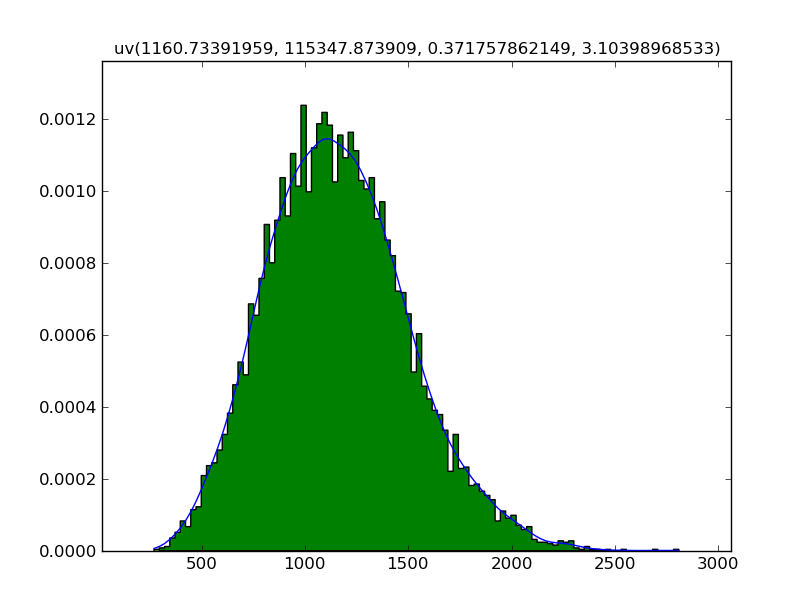
Since showing the plot is explicit, both views can be drawn together:
Z.plot()
Z.plot(hist=True)
Z.show()
Correlations¶
Use correlate before derived calculations when inputs should have a target
correlation structure:
import numpy as np
from mcerp import Exp, N, correlate, correlation_matrix, plotcorr
x1 = N(24, 1)
x2 = N(37, 4)
x3 = Exp(2)
correlation_matrix([x1, x2, x3])
[[ 1. 0.00558381 0.01268168]
[ 0.00558381 1. 0.00250815]
[ 0.01268168 0.00250815 1. ]]
The uncorrelated samples can be visualized with plotcorr:
plotcorr([x1, x2, x3], labels=["x1", "x2", "x3"], show=True)
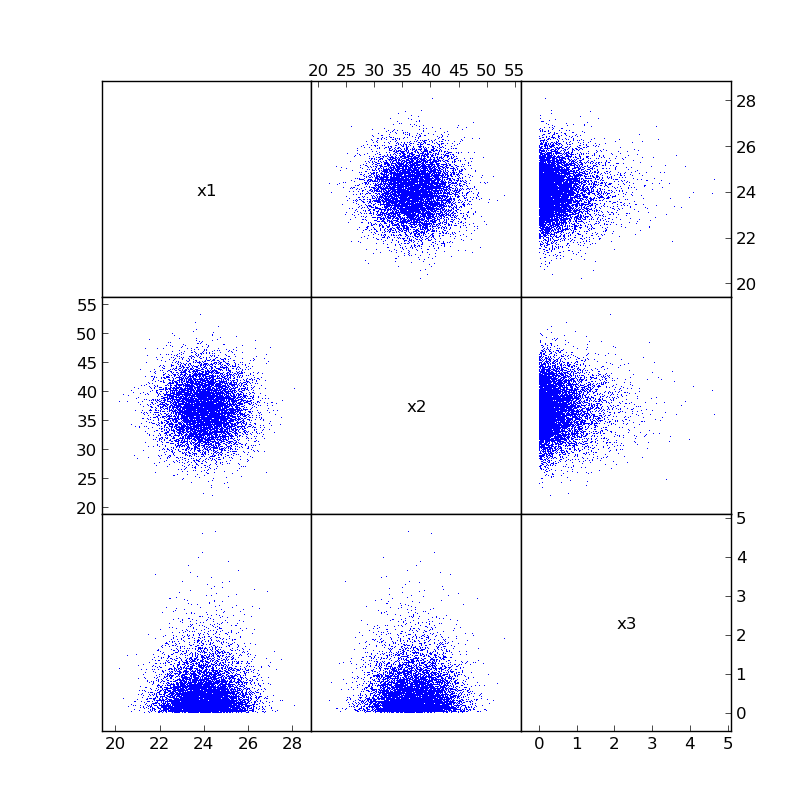
Now impose a target correlation matrix:
c = np.array([[1.0, -0.75, 0.0], [-0.75, 1.0, 0.0], [0.0, 0.0, 1.0]])
correlate([x1, x2, x3], c)
correlation_matrix([x1, x2, x3])
[[ 1.00000000e+00 -7.50010477e-01 1.87057576e-03]
[ -7.50010477e-01 1.00000000e+00 8.53061774e-04]
[ 1.87057576e-03 8.53061774e-04 1.00000000e+00]]
The newly correlated samples look like this:
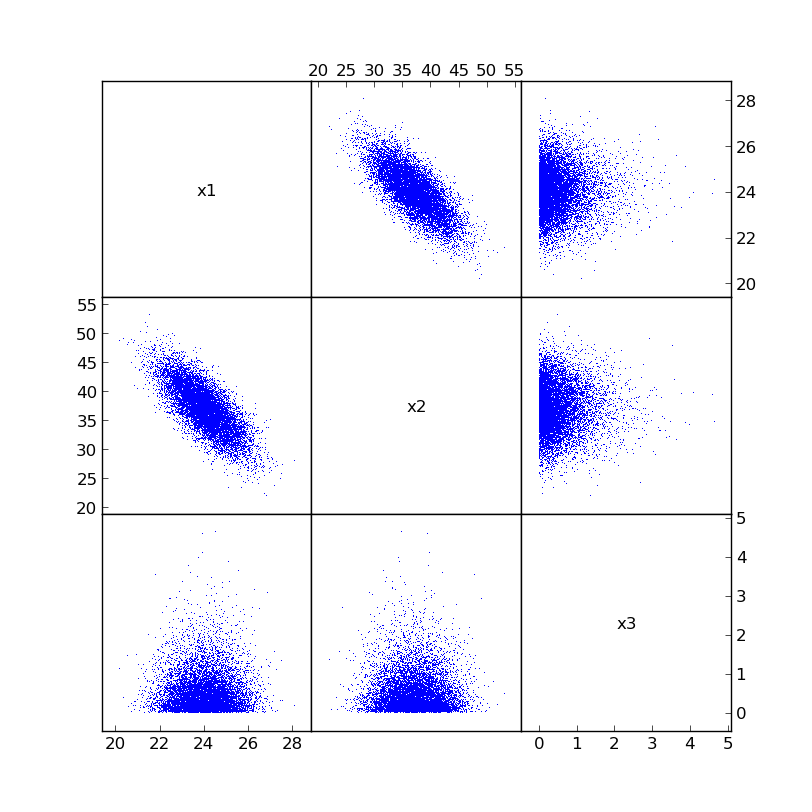
Now calculations based on x1, x2, and x3 use the reordered, correlated
sample points.
Z = (x1 * x2**2) / (15 * (1.5 + x3))
Z.describe()
MCERP Uncertain Value:
> Mean................... 1153.710442
> Variance............... 97123.3417748
> Skewness Coefficient... 0.211835225063
> Kurtosis Coefficient... 2.87618465139
The correlation operation does not change the original sampled values. It reorders them so that the inputs closely match the desired correlations.
Advanced Examples¶
Volumetric Gas Flow Through an Orifice Meter¶
This engineering example propagates uncertain measurements through a gas-flow calculation:
import mcerp.umath as umath
from mcerp import N
H = N(64, 0.5)
M = N(16, 0.1)
P = N(361, 2)
t = N(165, 0.5)
C = 38.4
Q = C * umath.sqrt((520 * H * P) / (M * (t + 460)))
Q.describe()
MCERP Uncertain Value:
> Mean................... 1330.9997362
> Variance............... 57.5497899824
> Skewness Coefficient... 0.0229295468388
> Kurtosis Coefficient... 2.99662898689
Even though the calculation involves multiplication, division, and a square
root, the result is close to a normal distribution: approximately
N(1331, 7.6).
Manufacturing Tolerance Stackup¶
Suppose a process fits a Gamma distribution with mean 1.5 and variance
0.25. Convert those statistics to Gamma shape parameters:
k = 1.5**2 / 0.25
theta = 0.25 / 1.5
x = Gamma(k, theta)
y = Gamma(k, theta)
z = Gamma(k, theta)
w = x + y + z
w.describe()
MCERP Uncertain Value:
> Mean................... 4.50000470462
> Variance............... 0.76376726781
> Skewness Coefficient... 0.368543723948
> Kurtosis Coefficient... 3.18692837067
Because the inputs are skewed, the output remains slightly skewed.
Scheduling Facilities¶
Six station cycle times can be combined into a full process-time distribution:
from mcerp import Chi2, Exp, Gamma, N
s1 = N(10, 1)
s2 = N(20, 2**0.5)
mn1 = 1.5
vr1 = 0.25
s3 = Gamma(mn1**2 / vr1, vr1 / mn1)
mn2 = 10
vr2 = 10
s4 = Gamma(mn2**2 / vr2, vr2 / mn2)
s5 = Exp(5)
s6 = Chi2(10)
T = s1 + s2 + s3 + s4 + s5 + s6
T.describe()
MCERP Uncertain Value:
> Mean................... 51.6999259156
> Variance............... 33.6983675299
> Skewness Coefficient... 0.520212339449
> Kurtosis Coefficient... 3.52754453865
The station standard deviations help identify where consistency matters most:
for i, si in enumerate([s1, s2, s3, s4, s5, s6]):
print("Station", i + 1, ":", si.std)
Station 1 : 0.9998880644
Station 2 : 1.41409415266
Station 3 : 0.499878358909
Station 4 : 3.16243741632
Station 5 : 0.199970343107
Station 6 : 4.47143708522
The probability of exceeding selected cycle times can be estimated directly:
[T > hr for hr in [59, 62, 68]]
[0.1091, 0.0497, 0.0083]